Blood cancer: Boosting chemotherapy effects
Lack of response to the DNA synthesis blocker cytarabine (ara-C) is the main cause for treatment failure and relapse.
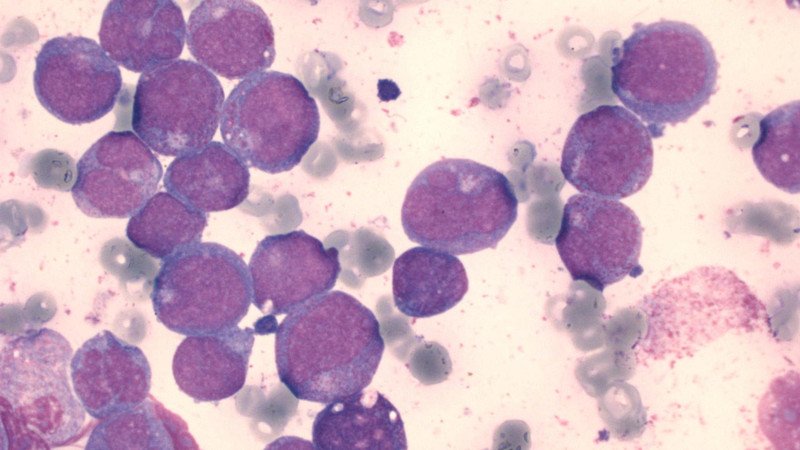
In December, a team headed by Jindrich Cinatl (Goethe-University Frankfurt) identified a new biomarker predicting poor response to the nucleotid analogue, which causes DNA damage through perturbation of DNA synthesis. Cytotoxicity of ara-C correlate with the ability of AML blasts (see photo) to accumulate the active metabolite ara-C triphosphate (ara-CTP). Cinatl’s team identified an enzyme that cuts off CTP, reconverting ara-C into its inactive form. According to the researchers, the CTPase code-named SAMHD1 could serve as a biomarker for responsiveness to cytarabine therapy.
In January, researchers headed by Thomas Helleday (Karolinska Institute, Stockholm) and Torsten Schaller (University Heidelberg) reported that blocking the SAMHD1 gene by CRISPR/Cas9-mediated gene disruption resensitised cancer cells to ara-C treatment. Furthermore, treatment with the Vpx protein of the simian immunodeficiency virus (SIV) dramatically increased sensitivity of primary patient-derived AML blasts to ara-C-induced cytotoxicity. In pediatric and adult AML patient cohorts receiving ara-C treatment, SAMHD1 presence proved to be a risk factor. The researchers thus conclude blocking SAMHD1 could become an attractive therapeutic strategy for boosting ara-C efficacy in blood cancers.